No unexpected CRISPR-Cas9 off-target activity revealed by trio sequencing of gene-edited mice | PLOS Genetics
Comparing -min-DP in vcftools with filter -i 'FORMAT/DP>10' in bcftools · Issue #1384 · samtools/bcftools · GitHub

Protocol for unbiased, consolidated variant calling from whole exome sequencing data - ScienceDirect

bcftools view | bcftools tutorial on how to count the number of snps and indels in a vcf file - YouTube
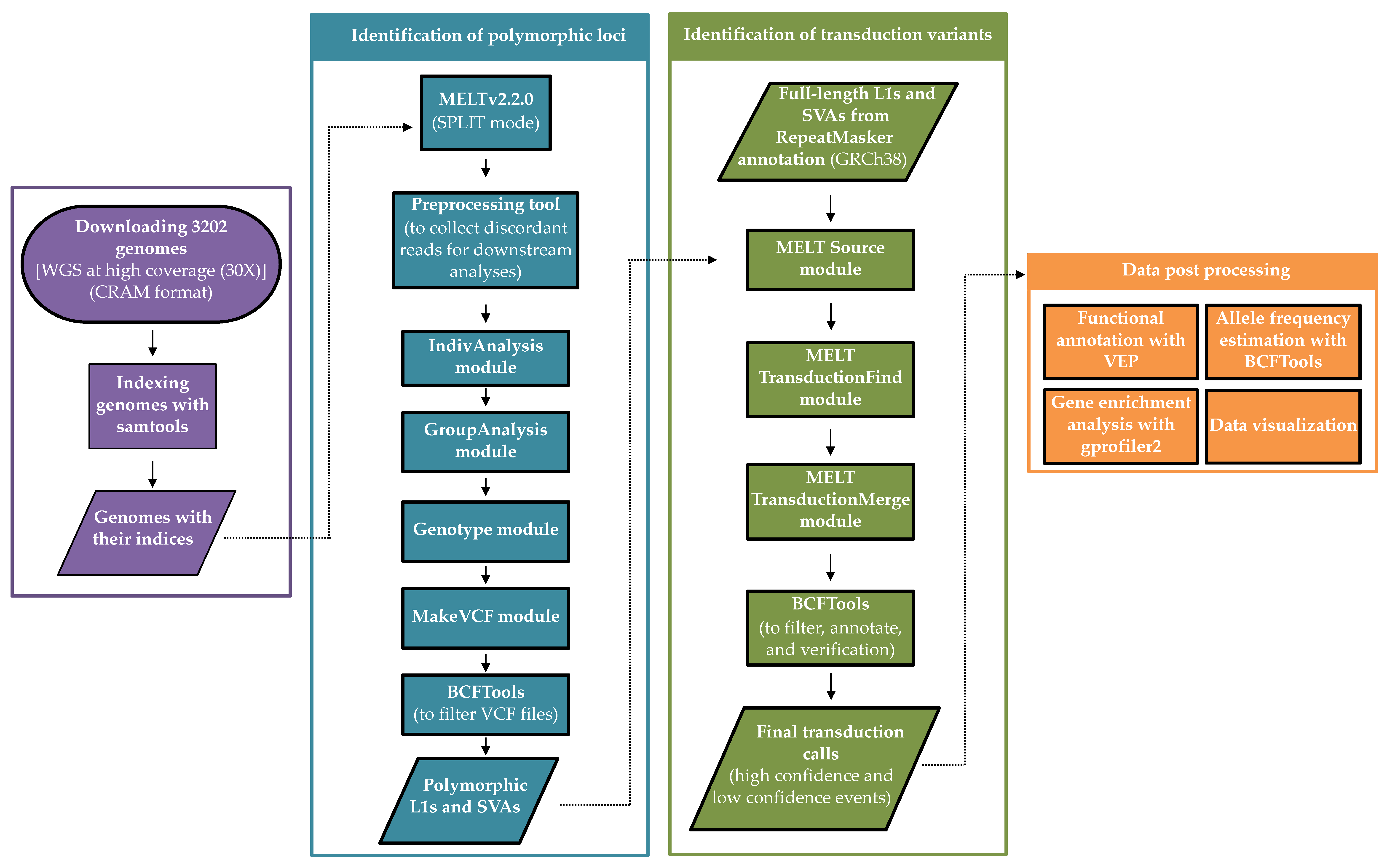
Biology | Free Full-Text | A Map of 3′ DNA Transduction Variants Mediated by Non-LTR Retroelements on 3202 Human Genomes

PDF) How to extract and filter SNP data from the genotyping-by- sequencing (GBS) data in vcf format using bcftools